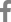Article Contents
| Clin Exp Pediatr > Volume 67(1); 2024 |
|
Bicaudal D is an important component of a crucial pathway that involves dynein and dynactin and is responsible for transporting mRNAs and other cellular cargo [1]. Variants in Bicaudal D2 Drosophila homolog 2 (BICD2) were known to be associated with autosomal dominant lower extremity-predominant spinal muscular atrophy 2 (SMALED2A/2B; Online Mendelian Inheritance in Man [OMIM]) [2,3]. After a recent study demonstrating loss of BICD2 function are associated with brain malformations in human [4], 9 patients with BICD2 variants were reported to be associated with brain malformations [5-10]. However, only a few patients have been reported to exhibit lissencephaly in brain imaging, and as such, this phenotype has not yet been added to the OMIM database. We report a case of Korean female patient with novel heterozygous BICD2 variant with developmental regression and lissencephaly, expanding neurologic phenotypic spectrum of BICD2-associated disease.
A 9-month-old female patient visited our outpatient clinic due to developmental delay. At the initial visit, she demonstrated eye contact, social smiles and head control, but was unable to sit unassisted. In neurological examination, spasticity and increased deep tendon reflexes were observed. Facial dysmorphism and joint abnormalities were not noted. Initial examination also revealed microcephaly, as head circumference measured 40.5 cm (-3.7 standard deviation). A brain magnetic resonance imaging taken at 9 months of age showed lissencephaly and corpus callosum hypogenesis (Fig. 1A). Routine laboratory tests, including creatine kinase, showed no notable abnormalities or neutropenia. During follow-up, she was able to sit alone at 14 months of age, crawl at 2 years of age, stand with assistance at 3 years of age, and understand a few commands. However, her developmental milestones regressed when she was 3.5 years old. The initial seizure occurred at 30 months of age, characterized by generalized seizures. A routine baseline electroencephalogram (EEG) at 16 months showed frequent repetitive beta activities, and the EEG at the onset of the first seizure revealed irregular high amplitude delta waves with a few suspicious frontal sharp waves. Initial treatment with valproic acid (up to 70 mg/kg/day) was followed by levetiracetam (up to 50 mg/kg/day) and topiramate (up to 50 mg/kg/day), but was still refractory. At 7 years old, the patient can only sit with assistance and does not understand commands. An EEG taken at 7 years old showed an excessive amount of high amplitude irregular delta waves and some suspicious frontal sharp waves. The patient still suffers from 2ŌĆō5 generalized seizures per day and did not gain any developmental milestones.
Trio whole exome sequencing (WES) was performed. The detailed method for data generation and processing was described elsewhere [11]. Initial trio WES analysis did not result in likely pathogenic/pathogenic variants linked to patientŌĆÖs phenotype, asBICD2 was reported to be associated with only spinal muscular atrophy phenotype and not with severe developmental delay and/or lissencephaly. A recent WES reanalysis revealed a novel de novo BICD2 variant (c.2324_2327dupAGAA [p.Thr777fs]) as a likely pathogenic variant with subsequent Sanger sequencing confirmation (Fig. 1B). All known BICD2 variants associated with brain malformations is shown in Fig. 1C.
Written informed consent-to-disclose was obtained from the parent of the patient. The study was approved by the Institutional Review Board (IRB) of Seoul National University Hospital (IRB No. 2010-125-1166).
BICD2 is vital for cerebral cortex development [4]. Only 9 patients with BICD2 variants were reported to have brain malformations (hypogenesis of corpus callosum, polymicrogyria, cerebellar hypoplasia, white matter loss and ventriculomegaly), including lissencephaly in 3 patients. Notably, microcephaly was observed in all BICD2-related lissencephaly cases unlike other BICD2-related brain malformations, while arthrogryposis commonly found in BICD2-related SMALED2A/2B patients was rare in lissencephaly patients. Our patient exhibited microcephaly and lissencephaly, consistent with previous studies [4,9,10]. We observed developmental regression in our patient, a novel occurrence in BICD2-related disease patients during long-term follow-up until age 7 (Table 1). We suspect it may result from refractory seizures not reported before.
We observed significant findings in understanding phenotype-genotype correlations of BICD2-related disorders. Most brain malformation-associated BICD2 variants occur in the conserved CC3 domain. Lissencephaly-related variants are typically heterozygous nonsense variants (p.Lys775Ter) in CC3 or homozygous variants (p.Leu244Pro and p.Gln77Ter) in CC1. Our patient's out-of-frame duplication variant in CC3 (p.Thr777fs) suggests that high-impact variants in CC3 and/or bi-allelic variants in other domains may contribute to the lissencephaly phenotype in BICD2-related disorders. No nonsense or out-of-frame deletion/duplication BICD2 variants associated with SMALED2A/2B phenotypes have been reported in the ClinVar database to date [12]. A recent study revealed interactions between the CC3 domain of BICD2 and 2 nuclear envelope-associated cargos, RanBP2 and Nesprin-2, in neuronal precursor cells in the developing brain. This finding suggests potential phenotype-genotype correlations with variants located in the CC3 domain [13].
In summary, we report a case of novel de novo heterozygous BICD2 variant-associated with lissencephaly and developmental regression. Given that lissencephaly phenotype has not yet been incorporated in the OMIM database, our case significantly contributes to the expanding phenotypic spectrum of BICD2-associated diseases and highlights the importance of raising awareness of the phenotypes reviewed in our study.
Fig.┬Ā1.
Radiogenomic findings of patients with BICD2-associated brain malformations in this study. (A) Brain magnetic resonance imaging findings of this patient with lissencephaly and hypogenesis of the corpus callosum (left: T1 sagittal; right: T2 axial). (B) PatientŌĆÖs pedigree and variant information. Trio-sanger sequencing was performed to confirm the variant. (C) All reported BICD2 variants associated with brain malformations to date. Variants associated with lissencephaly are marked in red.
Table┬Ā1.
Clinical phenotypes of patients with BICD2 variant-associated brain malformations
References
1. Hoogenraad CC, Akhmanova A, Howell SA, Dortland BR, De Zeeuw CI, Willemsen R, et al. Mammalian Golgi-associated Bicaudal-D2 functions in the dyneinŌĆōdynactin pathway by interacting with these complexes. EMBO J 2001;20:4041ŌĆō54.



2. Neveling K, Martinez-Carrera LA, H├Člker I, Heister A, Verrips A, Hosseini-Barkooie SM, et al. Mutations in BICD2, which encodes a golgin and important motor adaptor, cause congenital autosomal-dominant spinal muscular atrophy. Am J Hum Genet 2013;92:946ŌĆō54.



3. Kropatsch R, Schmidt HM, Buttkereit P, Epplen JT, Hoffjan S. BICD2 mutational analysis in hereditary spastic paraplegia and hereditary motor and sensory neuropathy. Muscle Nerve 2019;59:484ŌĆō6.



4. Tsai MH, Cheng HY, Nian FS, Liu C, Chao NH, Chiang KL, et al. Impairment in dynein-mediated nuclear translocation by BICD2 C-terminal truncation leads to neuronal migration defect and human brain malformation. Acta Neuropathol Commun 2020;8:106.




5. Fiorillo C, Moro F, Brisca G, Accogli A, Trucco F, Trovato R, et al. Beyond spinal muscular atrophy with lower extremity dominance: cerebellar hypoplasia associated with a novel mutation in BICD2. Eur J Neurol 2016;23:e19ŌĆō21.

6. Ravenscroft G, Di Donato N, Hahn G, Davis MR, Craven PD, Poke G, et al. Recurrent de novo BICD2 mutation associated with arthrogryposis multiplex congenita and bilateral perisylvian polymicrogyria. Neuromuscul Disord 2016;26:744ŌĆō8.


7. Koboldt DC, Waldrop MA, Wilson RK, Flanigan KM. The genotypic and phenotypic spectrum of BICD2 variants in spinal muscular atrophy. Ann Neurol 2020;87:487ŌĆō96.



8. Storbeck M, Horsberg Eriksen B, Unger A, Holker I, Aukrust I, Martinez-Carrera LA, et al. Phenotypic extremes of BICD2-opathies: from lethal, congenital muscular atrophy with arthrogryposis to asymptomatic with subclinical features. Eur J Hum Genet 2017;25:1040ŌĆō8.




9. Caglayan AO, Tuysuz B, Gul E, Alkaya DU, Yalcinkaya C, Gleeson JG, et al. Biallelic BICD2 variant is a novel candidate for Cohen-like syndrome. J Hum Genet 2022;67:553ŌĆō6.




10. Abdel-Salam GM, Girgis M, Eid MM, Sayed IS, Abdel-Hamid MS. A homozygous loss-of-function variant in BICD2 is associated with lissencephaly and cerebellar hypoplasia. J Hum Genet 2022;67:669ŌĆō73.




11. Park S, Jang SS, Lee S, Kim M, Sim H, Jeon H, et al. Systematic analysis of inheritance pattern determination in genes that cause rare neurodevelopmental diseases. Front Genet 2022;13:990015.








 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link PubMed
PubMed Download Citation
Download Citation